
Order the NovaSeq X Series
Advanced chemistry, optics, and informatics combine to deliver exceptional speed and data quality, outstanding throughput and scalability.
A fast, integrated workflow for preparing libraries for use in a wide range of sequencing applications.
Assay time
Hands-on time
Input quantity
Nextera DNA Flex products are now called Illumina DNA Prep. The (M) designation denotes a bead-linked transposome that generates an insert size of ~350bp. We recommend ordering the Illumina DNA/RNA UD Indexes Sets A-D (20091654, 20091656, 20091658, 20091660). The IDT for Illumina DNA/RNA UD Indexes Sets A-D (20027213, 20027214,20042666, 20042667) are being discontinued; the last order date is January 30th, 2025.
Illumina DNA Prep, previously known as Nextera DNA Flex, offers a fast, robust, and flexible workflow for preparing normalized, sequencing-ready libraries from a wide range of DNA input types and amounts facilitating an array of applications, from human whole-genome sequencing to sequencing amplicons, plasmids, and microbial species.1
Illumina DNA Prep workflow delivers consistent insert sizes, uniform coverage, and optimized performance, regardless of DNA input amount or genome size. The bead-based technology minimizes bias and opportunities for error, resulting in highly reproducible sequencing data.
The Illumina DNA/RNA UD Indexes Sets offer up to 384 unique dual indexes, enabling accurate assignment of reads and efficient use of the flow cell.
This product is also available as an Illumina Advantage (TG) product. Illumina Advantage large-scale sequencing products feature lot-specific shipments and testing, extended shelf life, and advanced change notifications for greater laboratory efficiency.
Find fast, optimized sequencing library preparation solutions for use in a wide range of applications.
A fast, integrated workflow for preparing libraries for use in a wide range of sequencing applications.
Illumina DNA Prep with Enrichment
A complete targeted resequencing solution for a wide range of applications, with fast integrated workflow and options for custom and fixed enrichment panels.
An efficient, fast, and integrated library preparation workflow that yields highly accurate data for sensitive sequencing applications.
| Assay time | ~3-4 hr (from DNA extraction to normalized library) |
|---|---|
| Automation capability | Liquid handling robot(s) |
| Automation details | Explore available automation methods |
| Description | A fast, flexible library prep workflow that accommodates an assortment of sample types and DNA input amounts, allowing access to a wide range of applications, from human to microbial whole-genome sequencing and more. |
| Hands-on time | 1-1.5 hr |
| Input quantity | 1-500 ng DNA for small genomes (e.g. microbial), or 100-500 ng DNA for large genomes (e.g. human). (For blood and saliva, see the reference guide). |
| Instruments | MiSeq System, iSeq 100 System, NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, MiSeqDx in Research Mode, MiniSeq System, NextSeq 550Dx in Research Mode, NovaSeq X System, NovaSeq 6000 System, NovaSeq X Plus System |
| Mechanism of action | Bead-linked transposome |
| Method | Amplicon sequencing, Whole-genome sequencing, De novo sequencing, Shotgun sequencing |
| Multiplexing | Up to 384 unique dual (UD) combinations and 96 combinatorial dual (CD) combinations |
| Nucleic acid type | DNA |
| Sample type details | Not validated for use with FFPE samples |
| Specialized sample types | Blood, Not FFPE-compatible, Saliva |
| Species category | Drosophila, Any species, Mammalian, Mouse, Yeast, Zebrafish, Human, Rat, Plant, Virus, Nematode, Bacteria |
| Species details | Compatible with any species |
| Target insert size | ~350 bp |
| Technology | Sequencing |
| Variant class | Single nucleotide polymorphisms (SNPs), Gene fusions, Loss of heterozygosity (LOH), Somatic variants, Chromosomal abnormalities, Germline variants, Structural variants, Insertions-deletions (indels), Copy number variants (CNVs) |
Each Illumina DNA Prep kit requires one of the index kits to complete the protocol, regardless of the sample pooling level used for sequencing. We encourage customers to use unique dual indexing whenever possible. Learn more about choosing the right indexing option for your application.
The index kits are NOT interchangeable with the Nextera DNA or Nextera XT Index Kits.
The Flex Lysis Kit is a separate product that complements the blood DNA extraction aspect of Illumina DNA Prep.
Illumina purification beads are included in the Illumina DNA Prep kit and do not need to be purchased separately.
Illumina DNA Prep supports a broad DNA input range (1–500 ng) and offers high-quality libraries for sequencing samples ranging from large whole genomes to amplicons, plasmids, and microbial species, in as little as ~3.5 hours.
Illumina DNA Prep
| Instrument | Recommended number of samples | Read length |
|---|---|---|
| NextSeq 550 System | 1 sample per run (high output; based on 30× coverage of a human genome) |
Up to 2 × 150 bp |
| NovaSeq 6000 System | 2–10 samples per run (dual flow cell; based on 30× coverage of a human genome) |
Up to 2 × 125 bp (rapid run) |
Microbial whole-genome sequencing
Use microbial whole-genome sequencing to map genomes of novel organisms, finish genomes of known organisms, or compare genomes across multiple samples.
Large whole-genome sequencing informs disease research and population genomics studies and reveals disease-associated alleles.
Comprehensively sample all genes in all organisms present in a given complex sample to evaluate bacterial diversity and detect unculturable microorganisms.
| Illumina DNA Prep | TruSeq DNA PCR-Free | TruSeq DNA Nano | Nextera XT DNA Library Preparation Kit | |
|---|---|---|---|---|
| Assay time | ~3-4 hr (from DNA extraction to normalized library) | 5 hr total assay time | ~6 hr total assay time | ~5.5 hr from DNA extraction to normalized library. (Library prep time: ~90 minutes). |
| Automation capability | Liquid handling robot(s) | Liquid handling robot(s) | Liquid handling robot(s) | Liquid handling robot(s) |
| Automation details | Explore available automation methods | Explore available automation methods | Explore available automation methods | Explore available automation methods |
| Description | A fast, flexible library prep workflow that accommodates an assortment of sample types and DNA input amounts, allowing access to a wide range of applications, from human to microbial whole-genome sequencing and more. | A simple, all-inclusive PCR-free prep for whole-genome sequencing studies with the ability to sequence through challenging regions of the genome. | A low-input research method that delivers high genome coverage quality and reduced bias. | Prepare sequencing libraries for small genomes, PCR amplicons, and plasmids in 90 min, with a low DNA input requirement. |
| Hands-on time | 1-1.5 hr | 4 hr | ~4 hr | 15 min |
| Input quantity | 1-500 ng DNA for small genomes (e.g. microbial), or 100-500 ng DNA for large genomes (e.g. human). (For blood and saliva, see the reference guide). | 1 ug DNA | 100 ng genomic DNA | 1 ng DNA |
| Instruments | MiSeq System, iSeq 100 System, NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, MiSeqDx in Research Mode, MiniSeq System, NextSeq 550Dx in Research Mode, NovaSeq X System, NovaSeq 6000 System, NovaSeq X Plus System | MiSeq System, NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, MiSeqDx in Research Mode, MiniSeq System, NovaSeq X System, NextSeq 500 System, NovaSeq 6000 System, NovaSeq X Plus System | MiSeq System, NextSeq 550 System, NextSeq 2000 System, MiSeqDx in Research Mode, MiniSeq System, NovaSeq X System, NextSeq 500 System, NovaSeq 6000 System, NovaSeq X Plus System | MiSeq System, iSeq 100 System, NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, MiniSeq System, NextSeq 500 System |
| Mechanism of action | Bead-linked transposome | Mechanical DNA fragmentation and adapter ligation. Workflow is PCR-free and gel-free. | Mechanical DNA fragmentation and adapter ligation. Workflow uses reduced-bias PCR and is gel-free. | Enzymatic fragmentation |
| Method | Amplicon sequencing, Whole-genome sequencing, De novo sequencing, Shotgun sequencing | Genotyping by sequencing, Whole-genome sequencing, Shotgun sequencing | Genotyping by sequencing, Whole-genome sequencing, Shotgun sequencing | Amplicon sequencing, Whole-genome sequencing, 16s rRNA sequencing, De novo sequencing, Shotgun sequencing |
| Multiplexing | Up to 384 unique dual (UD) combinations and 96 combinatorial dual (CD) combinations | Up to 24 single, 96 combinatorial (CD) dual, 24 unique dual and 96 unique dual (UD) combinations | Up to 24 single, 96 combinatorial (CD) dual, 24 unique dual and 96 unique dual (UD) combinations | Up to 384 uniquely indexed samples may be pooled and sequenced together. |
| Nucleic acid type | DNA | DNA | DNA | DNA |
| Sample type details | Not validated for use with FFPE samples | |||
| Specialized sample types | Blood, Not FFPE-compatible, Saliva | Not FFPE-compatible | Not FFPE-compatible, Low-input samples | Not FFPE-compatible, Low-input samples, Single cells |
| Species category | Drosophila, Any species, Mammalian, Mouse, Yeast, Zebrafish, Human, Rat, Plant, Virus, Nematode, Bacteria | Other, Mammalian, Mouse, Human, Rat, Plant | Other, Mammalian, Mouse, Human, Rat, Plant | Drosophila, Any species, Mammalian, Mouse, Yeast, Zebrafish, Fungal, Human, Rat, Plant, Virus, Nematode, Bacteria |
| Species details | Compatible with any species | Compatible with most large DNA genomes. | Compatible with most large DNA genomes. | Compatible with any species |
| Target insert size | ~350 bp | 350 bp or 550 bp | 350 bp or 550 bp | 300 bp–1.5 kb |
| Technology | Sequencing | Sequencing | Sequencing | Sequencing |
| Variant class | Single nucleotide polymorphisms (SNPs), Gene fusions, Loss of heterozygosity (LOH), Somatic variants, Chromosomal abnormalities, Germline variants, Structural variants, Insertions-deletions (indels), Copy number variants (CNVs) | Single nucleotide polymorphisms (SNPs), Gene fusions, Loss of heterozygosity (LOH), Somatic variants, Chromosomal abnormalities, Germline variants, Structural variants, Insertions-deletions (indels), Copy number variants (CNVs) | Single nucleotide polymorphisms (SNPs), Gene fusions, Loss of heterozygosity (LOH), Somatic variants, Chromosomal abnormalities, Germline variants, Structural variants, Insertions-deletions (indels), Copy number variants (CNVs) | Single nucleotide polymorphisms (SNPs), Structural variants |
Library Prep and Array Kit Selector
Find the right sequencing library preparation kit or microarray for your needs. Filter by method, species, and more. Compare, share, and order kits.
Sequencing Coverage Calculator
Calculator to help determine the reagents and sequencing runs needed to arrive at desired coverage for your experiment.
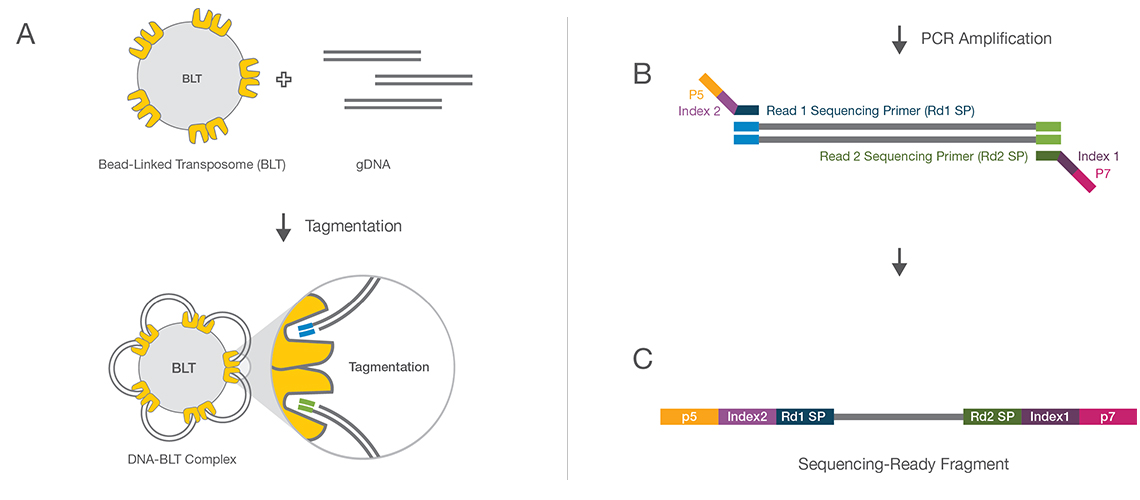

(A) Bead-linked transposomes mediate the simultaneous fragmentation of gDNA and the addition of Illumina sequencing primers. (B) Reduced-cycle PCR amplification amplifies sequencing ready DNA fragments and adds indexes and adapters. (C) Sequencing-ready fragments are washed and pooled.
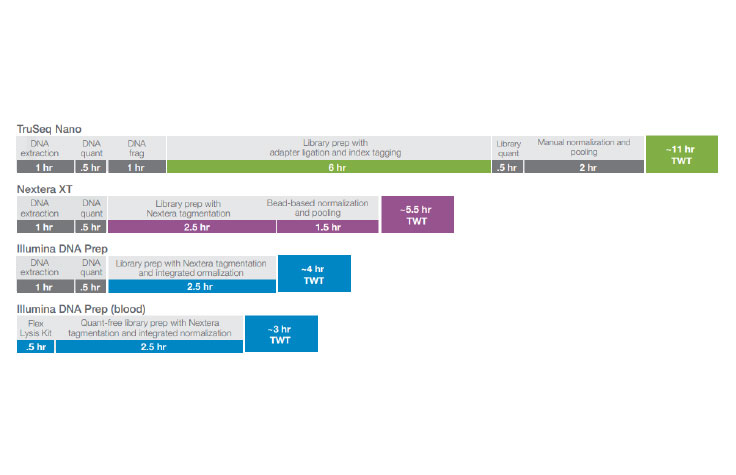

Comparison of the total workflow time from DNA extraction to library normalization and pooling. Illumina DNA Prep produces a workflow with the least number of steps and the fastest total workflow time.
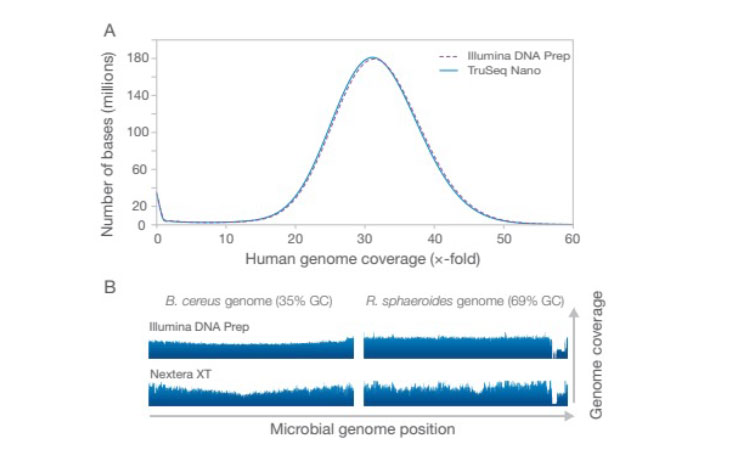

(A) Illumina DNA Prep delivers uniform coverage across the genome comparable to the TruSeq DNA Nano kit. (B) Coverage is shown for microorganisms with extremely high- or low-GC content. Due to improved on-bead tagmentation library prep chemistry, Illumina DNA Prep shows more even coverage than Nextera XT.

Learn about Illumina technologies designed to offer easy-to-use, fast, and simple solutions for both DNA and RNA library preparation.
On-bead tagmentation library prep chemistry integrates DNA extraction, fragmentation, library prep, and library normalization.
Illumina DNA Prep, (M) Tagmentation (24 Samples, IPB)
20060060
Includes reagents for preparing 24 libraries. Illumina Purification Beads are included. Purchase enrichment probe panel and index adapters separately.
List Price:
Discounts:
Illumina DNA Prep, (M) Tagmentation (96 Samples, IPB)
20060059
Includes reagents for preparing 96 libraries. Illumina Purification Beads are included. Purchase enrichment probe panel and index adapters separately.
List Price:
Discounts:
Illumina® DNA/RNA UD Indexes Set A, Tagmentation (96 Indexes, 96 Samples)
20091654
Illumina® DNA/RNA UD Indexes Set A, Tagmentation (96 Indexes, 96 Samples) Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
Illumina® DNA/RNA UD Indexes Set B, Tagmentation (96 Indexes, 96 Samples)
20091656
Illumina® DNA/RNA UD Indexes Set B, Tagmentation (96 Indexes, 96 Samples) Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
Illumina® DNA/RNA UD Indexes Set C, Tagmentation (96 Indexes, 96 Samples)
20091658
Illumina® DNA/RNA UD Indexes Set C, Tagmentation (96 Indexes, 96 Samples) Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
Illumina® DNA/RNA UD Indexes Set D, Tagmentation (96 Indexes, 96 Samples)
20091660
Illumina® DNA/RNA UD Indexes Set D, Tagmentation (96 Indexes, 96 Samples) Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
IDT® for Illumina® DNA/RNA UD Indexes Set A, Tagmentation (96 Indexes, 96 Samples)
20027213
Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
IDT® for Illumina® DNA/RNA UD Indexes Set B, Tagmentation (96 Indexes, 96 Samples)
20027214
Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
IDT® for Illumina® DNA/RNA UD Indexes Set C, Tagmentation (96 Indexes, 96 Samples)
20042666
Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
IDT® for Illumina® DNA/RNA UD Indexes Set D, Tagmentation (96 Indexes, 96 Samples)
20042667
Includes 96, 10 bp indexes sufficient for labeling 96 samples. Purchase library prep and enrichment reagents and probe panels separately.
List Price:
Discounts:
Nextera™ DNA CD Indexes (96 Indexes, 96 Samples)
20018708
Includes 20 indexes, eight of i500s and 12 of i700 indexes that together provide 96 dual indexing capability sufficient for labeling 96 samples. Indexes are provided in a 96 well format. Purchase library prep and probe panels separately.
List Price:
Discounts:
Illumina DNA Prep - Customer Site
20022900
A 1-day onsite training for the complete Illumina DNA Prep library preparation workflow. Participants will learn all steps of the workflow including sample input and library preparation, quality control, best practices, and troubleshooting, in addition to training on Illumina- supported sequencing and data analysis tools specific to the Illumina DNA Prep workflow.
Illumina Purification Bead, 100mL
20060057
Includes one, 100-ml container of beads for library clean-up and size selection. Purchase library prep and enrichment reagents, probe panels, and indexes separately.
List Price:
Discounts:
Illumina Purification Bead, 400mL
20060058
Includes four, 100-ml container of beads for library clean-up and size selection. Purchase library prep and enrichment reagents, probe panels, and indexes separately.
List Price:
Discounts:
Flex Lysis Reagent Kit (96 reactions)
20018706
Includes reagents for processing blood samples with the Illumina DNA Prep and Illumina DNA Prep with Enrichment. Purchase library prep, enrichment, enrichment probe panel, and index adapter reagents separately.
List Price:
Discounts:
Illumina® Free Adapter Blocking Reagent (12 Reactions)
20024144
This reagent is used during library preparation to reduce index hopping levels.
List Price:
Discounts:
Illumina® Free Adapter Blocking Reagent (48 Reactions)
20024145
This reagent is used during library preparation to reduce index hopping levels.
List Price:
Discounts:
NextSeq PhiX Control Kit
FC-110-3002
DNA control for the NextSeq Illumina sequencing platform. Compatible with single and paired-end reads up to 150 bp. 10 µl of a 10 nM PhiX solution.
List Price:
Discounts:
Showing of
Product
Qty
Unit price
Product
Catalog ID
Quantity
Unit price
Yes, Illumina has designed specific extraction protocols for both blood and saliva. The protocols can be found in the Illumina DNA Prep Reference Guide.
In general, Illumina DNA Prep is designed to be compatible with most automated liquid-handling systems.
All Illumina automation partners are developing methods for at least one existing automation platform. Contact the individual partners for more information.
Coverage of GC-rich regions can be impacted by the model, settings, and performance of the thermal cycler used. Illumina has validated the Bio-Rad DNA Engine Tetrad 2, the Bio-Rad S1000, the Bio-Rad C1000, and the MJ Research PTC-225 DNA Engine Tetrad thermal cyclers for use with Illumina DNA Prep. Other thermal cyclers may differ in their performance, which may impact genomic coverage.
Yes, the minimum input range (1–500 ng) of Illumina DNA Prep allows the use of DNA extracted from Dried Blood Spot (DBS) cards.
1. Bruinsma S, Burgess J, Schlingman D, et al. Bead-Linked Transposomes Enable a Normalization-Free Workflow for NGS Library Preparation. 2018;19:722. BMC Genomics. Published online 2018 Oct 1. doi: 10.1186/s12864-018-5096-9
Reach out for information about our products and services, or get answers to questions about our technology.
Your email address is never shared with third parties.